Fitting diagnostics: convergence, residuals, and uncertainty
Source:vignettes/fitting-diagnostics.Rmd
fitting-diagnostics.RmdOverview
fit_swrc() returns a convergence flag and standard
errors alongside the fitted parameters. This vignette shows how to:
- Check and handle non-converged fits
- Plot residuals to detect systematic model misfit
- Interpret parameter standard errors and compute confidence intervals
- Understand starting-value sensitivity, particularly for clay soils
1. Detecting non-converged fits
The convergence column is TRUE when
nls() converged and FALSE (with
NA parameters) when it did not.
set.seed(2947)
# Reference parameters for three soil types
soils <- tribble(
~soil, ~theta_r, ~theta_s, ~alpha, ~n,
"sandy loam", 0.047, 0.430, 0.145, 2.68,
"loam", 0.078, 0.430, 0.036, 1.56,
"silty clay loam", 0.089, 0.430, 0.010, 1.23 # difficult for NLS
)
heads <- c(0, 10, 33, 100, 330, 1000, 3300, 5000, 15000)
sigma <- 0.005
# Generate synthetic observations
obs <- soils |>
tidyr::crossing(h = heads) |>
mutate(
m = 1 - 1 / n,
theta_true = theta_r + (theta_s - theta_r) /
(1 + (alpha * h)^n)^m,
theta = pmax(0.01, theta_true + rnorm(n(), 0, sigma))
) |>
select(soil, h, theta)
fits <- obs |>
group_by(soil) |>
fit_swrc(theta_col = theta, h_col = h)
fits |>
select(soil, theta_r, theta_s, alpha, n, convergence)
#> # A tibble: 3 × 6
#> soil theta_r theta_s alpha n convergence
#> <chr> <dbl> <dbl> <dbl> <dbl> <lgl>
#> 1 loam 0.0731 0.429 0.0349 1.54 TRUE
#> 2 sandy loam 0.0469 0.431 0.141 2.71 TRUE
#> 3 silty clay loam 0.0298 0.427 0.00854 1.19 TRUE
n_failed <- sum(!fits$convergence)
cat(sprintf("%d / %d fits converged.\n", sum(fits$convergence), nrow(fits)))
#> 3 / 3 fits converged.
if (n_failed > 0) {
cat("\nNon-converged soils:\n")
fits |>
filter(!convergence) |>
pull(soil) |>
cat(sep = "\n")
}What to do when a fit fails:
- Try different starting values (see Section 4)
- Add more observations near the dry end (high |h|) where the curve flattens
- For very fine-textured soils (n < 1.2), consider fixing θ_s from bulk density measurements and fitting only θ_r, α, n
2. Residual analysis
A good fit should show residuals scattered randomly around zero. Systematic patterns suggest model misfit or bad observations.
# Reconstruct fitted curves and compute residuals
predicted <- fits |>
select(soil, theta_r, theta_s, alpha, n) |>
filter(!is.na(theta_r)) |>
tidyr::crossing(h = heads) |>
mutate(m = 1 - 1 / n) |>
swrc_van_genuchten(theta_r = theta_r, theta_s = theta_s,
alpha = alpha, n = n, h = h) |>
rename(theta_fitted = .theta)
resid_df <- obs |>
left_join(predicted |> select(soil, h, theta_fitted),
by = c("soil", "h")) |>
filter(!is.na(theta_fitted)) |>
mutate(residual = theta - theta_fitted)
ggplot(resid_df, aes(x = h, y = residual, colour = soil)) +
geom_hline(yintercept = 0, linetype = "dashed", colour = "grey60") +
geom_point(size = 2) +
scale_x_log10(labels = scales::label_comma()) +
facet_wrap(~soil) +
labs(
x = "Matric potential |h| (cm)",
y = expression(theta[obs] - theta[fit]),
title = "Residuals by soil type"
) +
theme(legend.position = "none")
#> Warning in scale_x_log10(labels = scales::label_comma()): log-10
#> transformation introduced infinite values.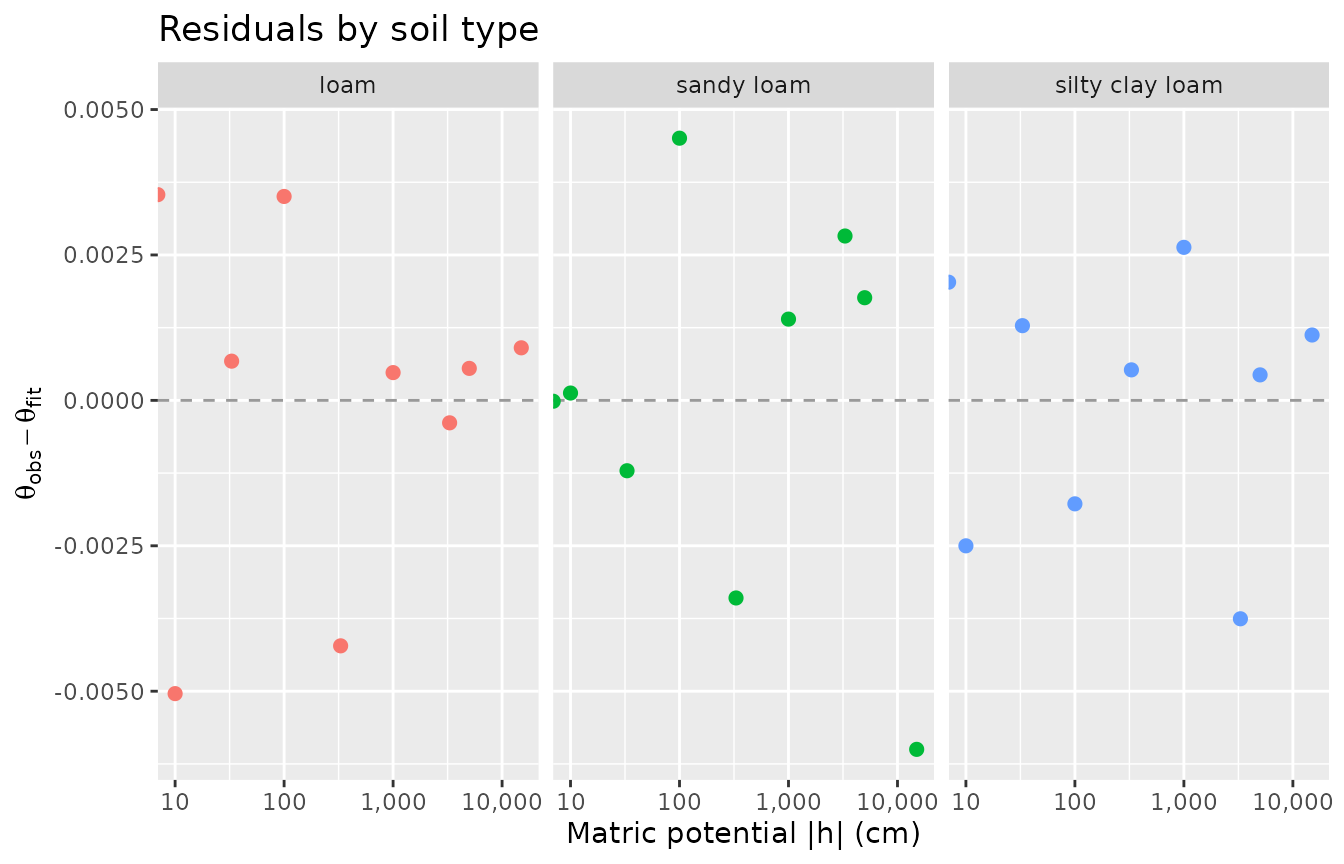
resid_df |>
group_by(soil) |>
summarise(
RMSE = sqrt(mean(residual^2)),
MAE = mean(abs(residual)),
max_e = max(abs(residual))
)
#> # A tibble: 3 × 4
#> soil RMSE MAE max_e
#> <chr> <dbl> <dbl> <dbl>
#> 1 loam 0.00279 0.00214 0.00504
#> 2 sandy loam 0.00303 0.00236 0.00600
#> 3 silty clay loam 0.00205 0.00179 0.003753. Parameter standard errors and confidence intervals
fit_swrc() returns standard errors for all four
parameters. For approximate 95% CIs use ± 2 SE; for exact intervals use
confint() on the nls object directly (see note
below).
fits |>
filter(convergence) |>
select(soil, theta_r, std_error_theta_r, alpha, std_error_alpha, n, std_error_n)
#> # A tibble: 3 × 7
#> soil theta_r std_error_theta_r alpha std_error_alpha n std_error_n
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 loam 0.0731 0.00450 0.0349 0.00264 1.54 0.0332
#> 2 sandy loam 0.0469 0.00176 0.141 0.00662 2.71 0.148
#> 3 silty cla… 0.0298 0.0464 0.00854 0.00118 1.19 0.0372
# 95 % CI: estimate ± 1.96 × SE
fits |>
filter(convergence) |>
mutate(
n_lo = n - 1.96 * std_error_n,
n_hi = n + 1.96 * std_error_n
) |>
select(soil, n, n_lo, n_hi, std_error_n)
#> # A tibble: 3 × 5
#> soil n n_lo n_hi std_error_n
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 loam 1.54 1.48 1.61 0.0332
#> 2 sandy loam 2.71 2.41 3.00 0.148
#> 3 silty clay loam 1.19 1.11 1.26 0.0372Interpretation:
- Narrow SE relative to estimate → parameter is well-constrained by the data
- SE larger than ~25% of estimate → consider adding observations at extremes of the retention curve (very wet or very dry)
- The
nparameter typically has the smallest relative SE when the data span a wide pressure range (several log-cycles)
4. Starting value sensitivity
Default starting values (θ_r = 0.9 × min(θ),
θ_s = 1.05 × max(θ), α = 0.1,
n = 2.0) work well for most mineral soils with n ≥ 1.4.
Clay soils (n ≈ 1.1–1.3) can require tighter, soil-specific starts.
# Difficult clay loam: n ≈ 1.23
clay_obs <- obs |> filter(soil == "silty clay loam")
# Strategy: grid search over (alpha, n) starting points
start_grid <- expand.grid(
alpha_start = c(0.005, 0.01, 0.02, 0.05),
n_start = c(1.1, 1.3, 1.5, 2.0)
)
fit_with_start <- function(a0, n0, obs_df) {
theta_vec <- obs_df$theta
h_vec <- abs(obs_df$h)
tryCatch({
fit <- nls(
theta ~ theta_r + (theta_s - theta_r) /
(1 + (alpha * h_vec)^n)^(1 - 1/n),
data = list(theta = theta_vec, h_vec = h_vec),
start = list(theta_r = min(theta_vec) * 0.9,
theta_s = max(theta_vec) * 1.05,
alpha = a0,
n = n0),
control = nls.control(maxiter = 200)
)
broom::tidy(fit) |>
tidyr::pivot_wider(names_from = term, values_from = estimate) |>
mutate(alpha_start = a0, n_start = n0, converged = TRUE)
},
error = function(e)
tibble(alpha_start = a0, n_start = n0, converged = FALSE)
)
}
start_results <- purrr::map2_dfr(
start_grid$alpha_start,
start_grid$n_start,
fit_with_start,
obs_df = clay_obs
)
start_results |>
select(alpha_start, n_start, converged, alpha, n) |>
arrange(desc(converged), n_start)
#> # A tibble: 52 × 5
#> alpha_start n_start converged alpha n
#> <dbl> <dbl> <lgl> <dbl> <dbl>
#> 1 0.005 1.1 TRUE NA NA
#> 2 0.005 1.1 TRUE NA NA
#> 3 0.005 1.1 TRUE 0.00854 NA
#> 4 0.005 1.1 TRUE NA 1.19
#> 5 0.01 1.1 TRUE NA NA
#> 6 0.01 1.1 TRUE NA NA
#> 7 0.01 1.1 TRUE 0.00854 NA
#> 8 0.01 1.1 TRUE NA 1.19
#> 9 0.005 1.3 TRUE NA NA
#> 10 0.005 1.3 TRUE NA NA
#> # ℹ 42 more rowsKey recommendations for clay soils
- α starting point: use 0.005–0.02 (reflecting fine-textured pores)
- n starting point: stay in the expected range 1.1–1.3 rather than defaulting to 2.0
- Data requirements: at minimum 8–10 observations spanning pF 0–4.2; include at least two points between pF 2–3 where clay SWRCs bend
- Fixed parameters: if K_sat is measured, pass it as θ_s constraint; reduces the fitting problem to three parameters
5. Visualising fitted curves alongside observations
# Only converged soils
fit_curves <- fits |>
filter(convergence) |>
select(soil, theta_r, theta_s, alpha, n) |>
tidyr::crossing(h = 10^seq(0, 5, by = 0.1)) |>
mutate(m = 1 - 1 / n) |>
swrc_van_genuchten(theta_r = theta_r, theta_s = theta_s,
alpha = alpha, n = n, h = h)
ggplot() +
geom_line(data = fit_curves,
aes(x = h, y = .theta, colour = soil),
linewidth = 0.9) +
geom_point(data = obs |> filter(soil != "silty clay loam"),
aes(x = h, y = theta, colour = soil),
size = 2.5) +
scale_x_log10(labels = scales::label_comma()) +
labs(
x = "Matric potential |h| (cm)",
y = expression(theta~(m^3/m^3)),
colour = "Soil",
title = "Observed (points) vs fitted VG curves"
)
#> Warning in scale_x_log10(labels = scales::label_comma()): log-10
#> transformation introduced infinite values.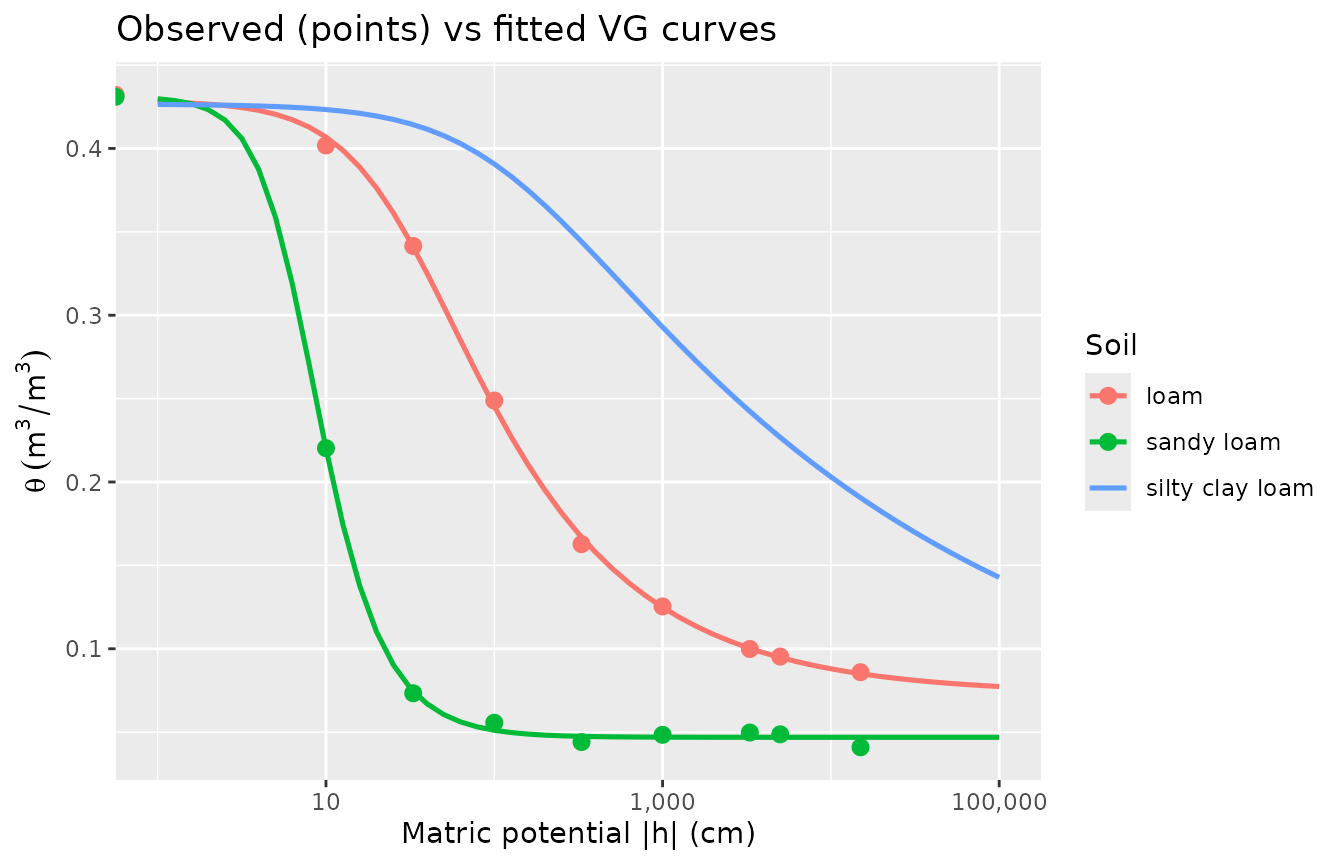
Summary
| Diagnostic | What to check |
|---|---|
convergence column |
FALSE → adjust starting values or data range |
| Residual plot | Systematic patterns → model misfit; outliers → bad observations |
std_error_* columns |
SE / estimate > 0.25 → more data needed |
| Starting value grid | Clay soils: try α ∈ [0.005, 0.02], n ∈ [1.1, 1.3] |